At the 2026 Strasbourg Interop
It's Interop time again!

Semiannually, the VO community meets to discuss what we've done since the last Interop and what needs to be done in the future. This week it does so again, this time in Strasbourg (programme).
The posts tagged with Interop will give you an impression of how these meetings feel, and I'd like to do some close-to-real-time blogging about this one again; just come back here occansionally until Friday if you are curious.
Right now, I am sitting in the opening session, remembering how, half a year ago, I was hectically trying to keep everything together at the Görlitz Interop when I was the local organising committee. Oh, how much more professional everything is here in Strasbourg's manufacture des tabacs: Good sound, zoom room working on the first attempt, no sun rays blotting out the projection screen, eduroam internet. It's really a nice lecture hall here, where we had to cobble together something rather improvised in Görlitz. Infrastructure matters.
Memories aside, the first talk of an Interop traditionally is the State of the IVOA delivered by the chair of the Exec, which quite as traditionally sports slides from the member organisations. I had to smile and couldn't help being flattered when JJ took up my nerd theme and quipped about “what the nerds like“ or so on GAVO's slide.
Going on, I can't resist a piece of trivia: The claim of the report on the activities of the Committee for Science Priorities was “no acronyms“ (and I will give you that for outsiders, the density of odd words between ADQL and UWS is a bit scary). Well: It was 22. A colleague counted. But then by Interop standards, that's still a pretty impressive achievement.
Oh, and the State of the TCG closed a whopping seven minutes before then end of the session, and now a discussion on how to do VO propaganda has ensued. People make a point that there's few things more useful for that than hands-on courses. Which has been a cue for me because since the last Interop, my pet project DocRegExt is exactly designed for that and became an official standard (“recommendation“) just in the last semester. Ha!
Monday 15:00 – The Local Host Session
The first “business“ session of this Interop has talks advertising the achivements of the “local“ VO enthusiasts, where at first “local” means French.
Ada Nebot's talk on OV [sic!] France is a bit humbling for me. For instance, they have a mailing list for technical discussions with more than 100 subscribers – wow. In Germany, at GAVO, we never made it beyond a dozen for our equivalent. Perhaps I should have worked a bit harder on hauling in money after all?
But then of course France profits from a far-sighted personnel planning: There actually have been permanent positions for data curation and publication over here since several decades. Let's see how this pans out back in Germany – this year, we will fill the first positions that are at least planned to become permanent for the new data centre at the DZA in Görlitz.
Carolin Bot then relates stories about the CDS, which I'd chalk down as the most important data centre in the VO, partly because of the Simbad database. This, I just learned from Carolin, collects object data from a whopping 15'000 articles per year.
I get queasy when I consider that there are close to 100 new scientific articles in Astronomy alone every working day that Simbad processes (which means that there's a lot more that don't talk about objects). I can't resist mentioning that we really need to fix our publication system by either getting rid of performance-fantasising metrics altogether (which would be my preferred outcome) or at least use something else than publications. Still, great work, Simbad. Thanks, and thanks a lot for your TAP service, which is an incredibly powerful tool. If you, dear reader, do not know what I'm talking about, by all means check out our VO course (which features it).
Talking about metrics abuse: Carolin also reports the CDS is serving 5 million queries per day. I'd certainly not want to use this as a proxy for CDS' usefulness – one smart TAP query or a catalogue crossmatch could replace a million requests each while providing a better service –, but it means that CDS' servers have to withstand 50 requests per second on average. Even if modern computers are amazingly fast, that is a certain challenge, in particular considerung that some of these requests can cause many seconds of computation.
Hours, actually, if you don't pay attention to efficiency. Fortunately for CDS, there are people there who actually look at efficiency and realise there's a difference between code that takes half a second on the one hand and code that takes 50 ms on the other hand – something I myself rarely indulge in.
And then there was a great slide in Andy Götz' concluding talk on the European Open Science Cloud EOSC (the “local” is Europe in that case), where he makes it clear that the EOSC is not a cloud, not (only) European (because “open” only makes sense if you don't close out the rest of the world), and regrettably it's not always open, either. If there is a useful definition of the EOSC beyond “a funding scheme of the EU“, however, I still could not figure out.
But then I freely confess to being very skeptical about discipline-spanning data publication in the first place on grounds that there are not many problems that the different disciplines actually share; I couldn't name much beyond AAI (i.e., authentication, which I'd rather not have at all in the first place) and PIDs (persistent identifiers; and these are a lot less useful on top of non-permanent infrastructure than you would think). Let me stress that I'm saying this as someone who's been soliciting contributions to my Stories on Cross-Discipline Data Discovery for a long time. You'd be surprised how little enthusiasm for this kind of thing is out there.
Monday 17:00 – Apps I
I'm sitting in the first session of the Appliations Working Group (“Apps”), which in VO circles is affectionally known as Show & Tell.
Against this cliche, the first talk (sorry, no link: it would go to Google docs) is about HATS, a fairly cool new format for dealing with large catalogues without having to run TAP and ADQL. It is a bit of a cross between HiPS and Parquet. By Apps standards, though, the talk it was fairly technical and had few colourful pictures. You could argue that is a quality, and I could not deny that.
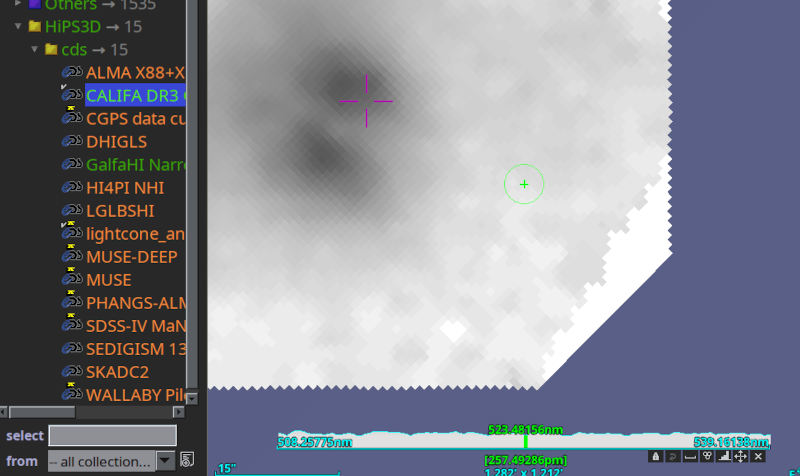
Things became a bit more baroque in the next talk: Pierre Fernique had the chair turn the light down before starting his slides – and will, I think, now show how you can interact with a data cube of 600 GB (a) at all, (b) over the network, and (c) on a very moderate machine. This already works from the comfort of your home (or office) with the most recent Aladin beta (v12.675). Try the HIPS3D subtree in the discovery tree; lightcone is fun, but being able to zoom through the spectral cubes from CALIFA to me is more impressive:
FX Pineau next reported on HATS progress (among other things), namely that the CDS now produces the HATS files I talked about above on the fly. Hu. Should DaCHS know how to do that, too?
Beyond that, in that talk you can see a few instances of what I was referring to above when I said CDS folks do consider efficiency. I like it if even today software people still consider the number of disk seeks required to do what their programs try to do. Yes, I know that with SSDs they're not nearly as expensive any more as they used to be, but, you know, my mass data is also still served from spinning disks.
Tuesday 10:00 – Sciene Platforms
At this Interop, there is a plenary session on science platforms. I have already ranted about this return of the data silos during the last interop. In Tess' talk, there was a slide that nicely summarises my concerns:
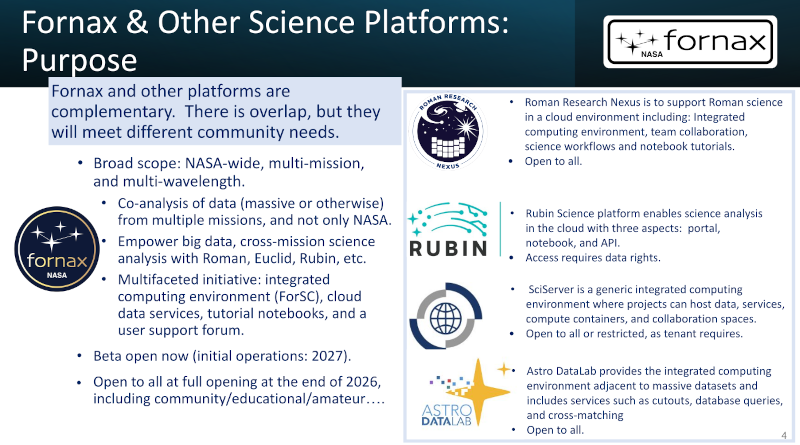

So: everyone spends a lot of effort on building complex systems of their own that can (in the most extreme case) process just a single sort of data (theirs), requiring different credentials, different code, that cannot interoperate, that are pretty much silos that, when they go down, will take all the software and workflows written for them with them. Most of them, I think, also depend on AWS, and what happens when Amazon changes their rules and/or pricing is anybody's guess.
Tuesday, 12:00 – DAL I
In the first DAL session of this Interop, I'm a bit distracted because back in Heidelberg our computation centre has again cut off our servers. After they had a two day “power outage“ over pentecost and the still-unresolved November disaster, I again regret the day when the University forced us to move our servers to them.
I still appreciated Pat's remark in his talk on OpenAPI for TAP 1.2 that if we could go back in time, we certainly wouldn't make our protocols' query parameter names case-insensitive. Absolutely. I'd widen that statement: Whenever you think that case-insensitivity is a good idea, you are probably wrong. My experience is that this will almost certainly going to come back at you later to no end of headache. Just have a look at the RegTAP spec and search for “case”. Each of these places cost me a bunch of hair.
Case folding considered harmful. Let's not do it any more.
I was re-inforcing that point in the first talk I was giving at this Interop, SCS-2.0 prototype implementation
In there, I reported that purging case-folding from some parameters of SCS2 (the one it shares with case-folding protocols) was really painful – but also unavoidable. Other than that, I was delighted that there was a lively discussion afterwards; at least there's interest in the activity, albeit it seems more in the protocol itself rather than the management of a major version transition that is, really, why I am after SCS2.
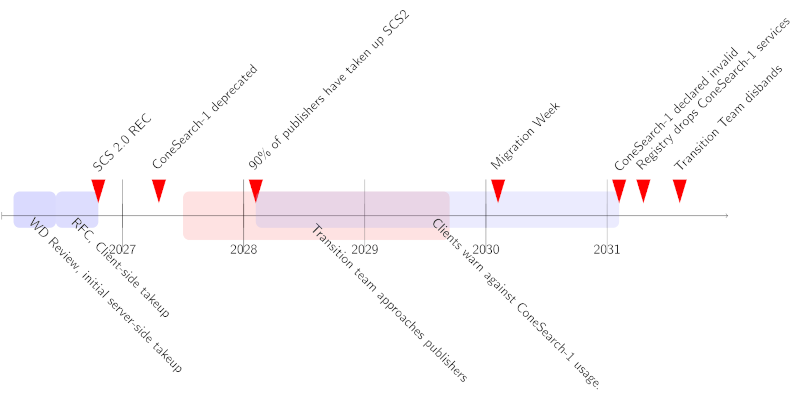
It would seem, anyway, that we will go on with SCS2. There's a time line in the SCS2 draft that covers something like five years. In that sense, this session may very well have been the point of no return for a long, long journey.

![[RSS]](./theme/image/rss.png)